
电话:+86-755-86967710
邮箱:webmaster@szbl.ac.cn

Institute of Systems and Physical Biology
zhouyq@szbl.ac.cn
Computational Biology,Molecular Biology ,Microbiology
Senior Principal Investigator
Professor
Tenured Full Professor
Tenured Associate Professor
Assistant Professor
Research Associate
Postdoctoral Fellow
Postdoctoral Fellow
Scientist and Director
The research group led by Dr. Zhou primarily focuses on fundamental studies exploring the relationships between sequences, structures, and functions of RNA and proteins, as well as applied research in design, delivery, and drug development of these biological macromolecules. A distinctive feature of the group is the integration of dry and wet lab approaches—combining computational structural bioinformatics, AI-powered deep learning with modern high-throughput and automated directed-evolution biotechnologies—to achieve profound insights into the sequence-structure-function relationships. This research enables the multi-faceted application of biological macromolecules, including targeted drug design and delivery for precision medicine, as well as detection of personalized biomarkers. Current specific projects include, but are not limited to:
1. Protein structure and function prediction in the post-AlphaFold era
2. Protein design and directed evolution
3. RNA structure prediction and RNA language models
4. Targeted RNA delivery systems
Pioneered prediction of continuous backbone dihedral angles using shallow and deep learning, enabling end-to-end protein structure prediction (Fig. 1). This development directly underpinned Nobel-prize-winning technique AlphaFold 2 for high-accuracy structure prediction.
First AI-driven methods for protein sequence design, achieving 30–34% sequence recovery (Fig. 2). Recognized as the inception of machine learning in protein design ([Ovchinnikov and Huang, Curr. Opin. Chem. Biol. 2021]), this paradigm shift (away from energy-based approaches) now dominates the field, enabling revolutionary advances in therapeutic protein & industrial enzyme design.
RNA-BRiQ: Developed a novel statistical potential enabling atomic-level RNA refinement. Powered AIChemy-RNA2 to win #1 in CASP15 (2022) for RNA structure prediction.
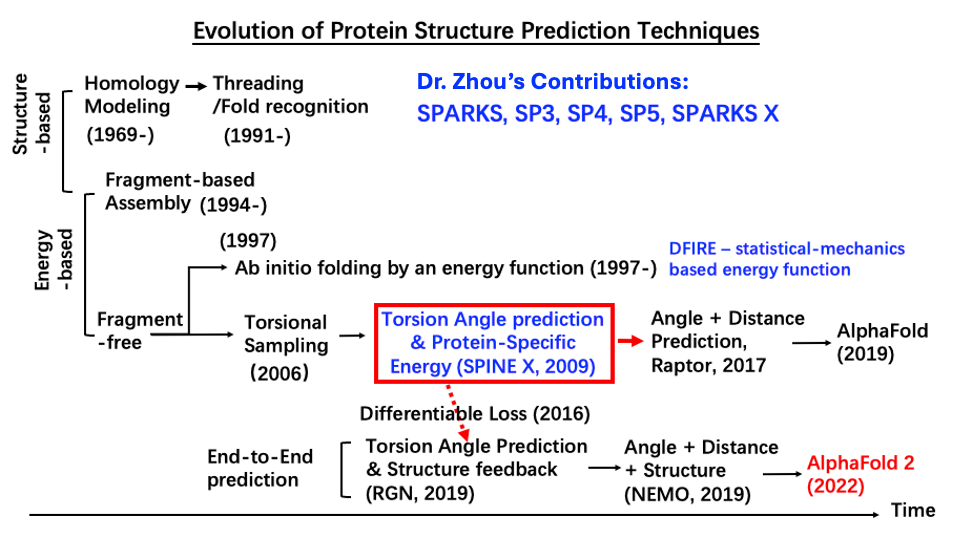
(Fig. 1)
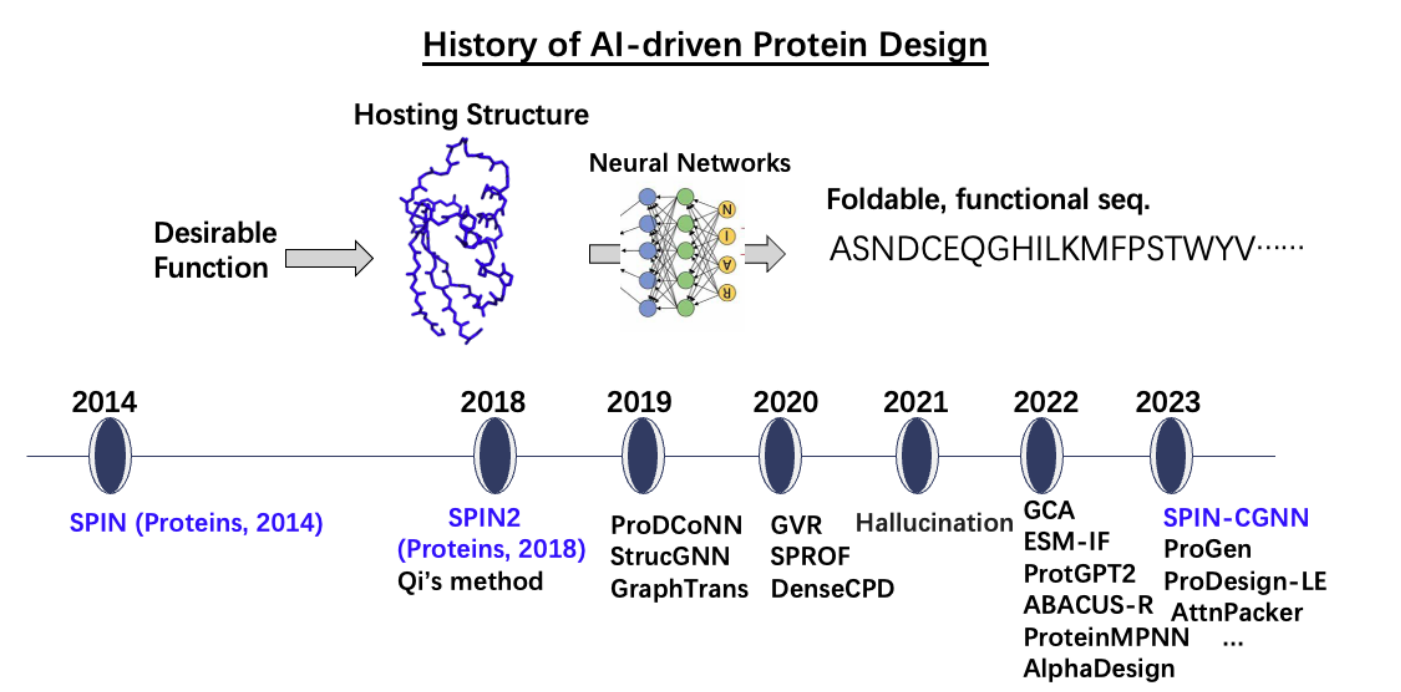
(Fig. 2)
1. J. Singh, J. Hanson, K. Paliwal, and Y. Zhou, RNA secondary structure prediction using an ensemble of two-dimensional deep neural networks and transfer learning,Nature Communications10, 5407 (2019).
2. Z. Zhang, P. Xiong, T. Zhang, J. Wang, J. Zhan, and Y. Zhou, Accurate inference of the full base-pairing structure of RNA by deep mutational scanning and covariation-induced deviation of activity,Nucleic Acids Research, 48:1451-1465 (2020).
3. J. Zhan, H. Jia, E. A. Semchenko, Y. Bian, A. M. Zhou, Z. Li, Y. Yang, J. Wang, S. Sarkar, M. Totsika, H. Blanchard, F. E.-C. Jen, Q. Ye, T. Haselhorst, M. P. Jennings, K. L. Seib, and Y. Zhou, Self-derived structure-disrupting peptides targeting methionine aminopeptidase in pathogenic bacteria; a new strategy to generate antimicrobial peptides,FASEB J., 33: 2095–2104 (2019).
4. S. Xu, J. Zhan, B. Man, S. Jiang, W. Yue, S. Gao, C. Guo, H. Liu, Z. Li, J. Wang, and Y. Zhou, Real-time reliable determination of binding kinetics of DNA hybridization using a multi-channel graphene biosensor,Nature Communications8, 14902 (2017).
5. Z. Li, Y. Yang, J. Zhan, L. Dai and Y. Zhou, Energy Functions in De Novo Protein Design: Current Challenges and Future Prospects,Ann. Rev. Biophysics42, 315-335 (2013).
Contact
Address: Gaoke Innovation Center,Guangming District, Shenzhen
Phone: +86-755-86967710
Email: webmaster@szbl.ac.cn
Postal Code: 518132
Copyright © 2025 Shenzhen Bay Laboratory. All Rights Reserved.