Prof. SUN Kun’s group from SZBL recently published their latest research on Bioinformatics, a leading journal in the field of computational biology. Their research deliverable, Ktrim, is a newly developed software that serves as a valuable and efficient tool for short-read sequencing data preprocessing which helps remove the adapter sequences and low quality cycles to facilitate downstream analyses of sequencing data.
1. What is a sequencing adapter?
Next-Generation Sequencing (NGS) is a widely used technology in biomedical studies. DNA of interests must go through a series of chemical treatments in order to be compatiable with the sequencers, which include a step that adds adapter sequences to the DNA molecules. The adapters are designed to align the DNA of interests to the flowcells and make it recognizable to the sequencing primers.
2. Why the adapters must be removed during data analysis?
The DNA sequencer read the DNA molecules of interest from one side to the other. When the sequencing read length is longer than the DNA molecule, the adapter sequence will be encountered and reported in the final sequencing reads. DNA reads that are contaminated by the adapter sequences (and low quality cycles which usually appear at the tail) cannot be aligned to the reference genome, and also introduce artefacts in downstream analyses (e.g., somatic mutation calling) As a result, the adapter sequencing step has become a universal preprocessing process in NGS data analysis.
3. What are the advantages of Ktrim?
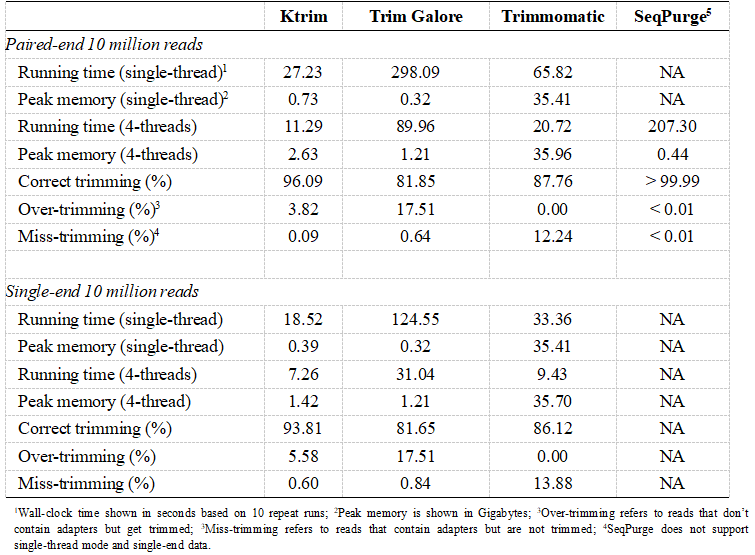
With the ever-growing throughput and read length of modern sequencers, the preprocessing step turns to be a bottleneck in data analysis due to unmet performance of current tools. Extra-fast and accurate adapter- and quality-trimming tools for sequencing data preprocessing are therefore still of urgent demand. To this end, Prof. SUN’s group developed the Ktrim software, which provides much stronger performance along with high accuracy and balanced sensitivity and specificity. Ktrim has been benchmarked against current tools including Trim Galore(2011; based on cutadapt), Trimmomatic(2014) and SeqPurge(2016). Ktrim was ~2-18 times faster than current tools and also showed high accuracy (Table 1). Moreover, the performance of Ktrim was barely unaffected when dealing with sequencing reads with an error rate as high as 5%. Ktrim is also a versatile tool: it provides built-in support to adapters of common library preparation kits; supports user-supplied, customized adapter sequences; supports both paired-end and single-end data; supports parallelization to accelerate the analysis. Ktrim could thus serve as a valuable and efficient tool for short-read NGS data preprocessing in the big data era.
Table: Performance comparison of Ktrim and current tools
PubMed:https://www.ncbi.nlm.nih.gov/pubmed/32159761
DOI:https://doi.org/10.1093/bioinformatics/btaa171
Ktrim Download:https://github.com/hellosunking/Ktrim
