Recently,MindSpore Sponge, a molecular simulation library was launched. Jointly developed by SZBL, Gao Yinqin research group of PKU and Huawei MindSpore, Sponge provides high performance, modularity and related support services for research demanding quick processing of both theory and algorithm, and facilitates natural fusion of artificial intelligence and molecular simulation.
Being a kind of computer simulation method, molecular dynamics simulation leverages Newton's law to describe the micro evolution of atoms and molecules. Molecular dynamics simulation can be applied to basic research and industrial senario. It is conducive for scientific researchers in the basic research community to study the physicochemical property of the system at microscopic level.
For the industrial sector, molecular dynamic simulation supports drug molecules design and the search of protein targets with large scale computing capability. Traditional molecule dynamics software experiences a rather narrow scope in actual application due to the limitation of simulation time and space. As an effort to widen the application scope, researchers have been working to find possible solutions like developing new force field models and sampling methods, or combining AI technology with these software.
To this end, the development of a new generation of molecular dynamic software, which is modular and highly-efficient for building structures that can verify theoretical models, is of great necessity. More than this, the software also needs to have the compatibility for application under traditional senario, be able to realize natural fusion of molecular simulation plus machine learning, and possess the form with embedded AI frameworks.These ideas explain how Sponge, the fully automated software, were created.
Sponge offers higher efficiency for using SITS to simulate biosystem through offering native support for SITS and optimized calculation processes, compared with enhanced sampling delivered via combination of traditional molecular simulation software and SITS methods. Traditional molecular simulation usually combines quantitative calculation to cope with charge fluctuation in polarized systems. However, even if machine learning algorithm is used to achieve calculation reduction, a great amount of time will still be needed for data transmit. The good news is that Sponge is able to shorten the calculation time by using modularity features to support the direct communication between the memory storage and the machine learning program.
Figure 1: SITS enhanced-sampling molecular dynamics simulation on alanine dipeptide in soluble developer
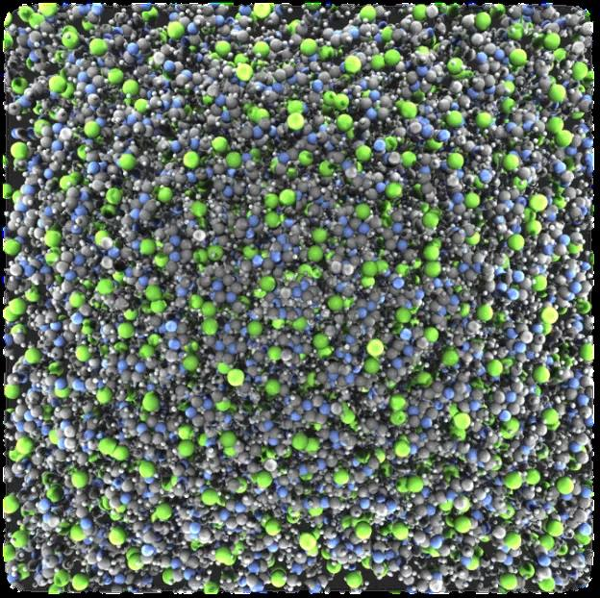

Figure 2: Machine learning + molecular simulation provide quick simulation on polarized system The graph is [C1MIm]Cl Ionic liquid simulation
Sponge can perform and complete traditional molecular simulation under high efficiency based on MindSpore automatic parallelization and image fusion algorithm. Drawing on MindSpore automatic differentiation, Sponge can combine traditional molecular simulation with neural networks and related AI approaches.
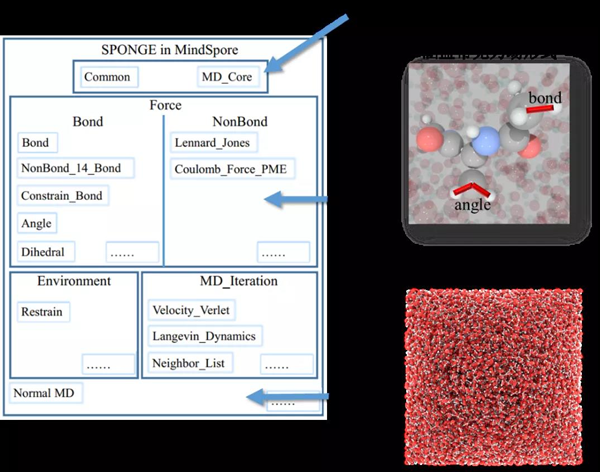

Figure 3: Sponge modular design structure
MindSpore + Sponge | The Advantages of Sponge
1. Full modularity in molecular simulation. Sponge provides modularized construction of molecular simulation algorithms and favorable open source community for the external to share sub modules.
2. Full process of AI algorithm of traditional molecular simulation combining MindSpore. MindSpore makes it easier for researchers to apply AI methods in molecular simulation. Fully operatorized Sponge and MindSpore will be further integrated into a new generation of end-to-end differentiable molecular simulation software to achieve natural fusion of AI and molecular simulation.
Tutorial
https://www.mindspore.cn/tutorial/training/zh-CN/r1.2/advanced_use/hpc_sponge.html
Outlook
Near-termupdate of Sponge will be added with theoretically-verified MetalTS module, finite element calculation module and etc. These modules will improve Sponge’s performance in phase transition and metal surface simulation. Also, MindSpore Sponge will assist automatic differentiation and parallelization, providing more user- friendly support to cooperate with machine learning program.
Long-termedition of Sponge will expand its featured modules to enable description on most micro systems and improve calculation and sampling efficiencies. As for specific industrial demands, drug screening for example, a comprehensive calculation scheme will be derived based on Sponge to meet the demands of large scale parallel computation. Under the MindSpore framework, Sponge will possess meta optimization function to achieve force field fitting with higher accuracy and speed.
